Multilevel models
Professor Andy Field
University of Sussex





|
Learning outcomes
- Understand what hierarchical data are
- Why we can’t use the OLS GLM
- Understand fixed and random coefficients
- Understand how to build models
- Be able to conduct and interpret models of hierarchical data
Hierarchical data
- Data structures are often hierarchical
- Children nested within classrooms
- Observations nested within people
- Employees nested within organisations
- Patients nested within hospitals
- Patients nested within teams nested within hospitals
- Service users nested within clinicians nested within hospitals nested within NHS trusts!
- Zombies nested within rehabilitation clinics 😉
A two-level hierarchy
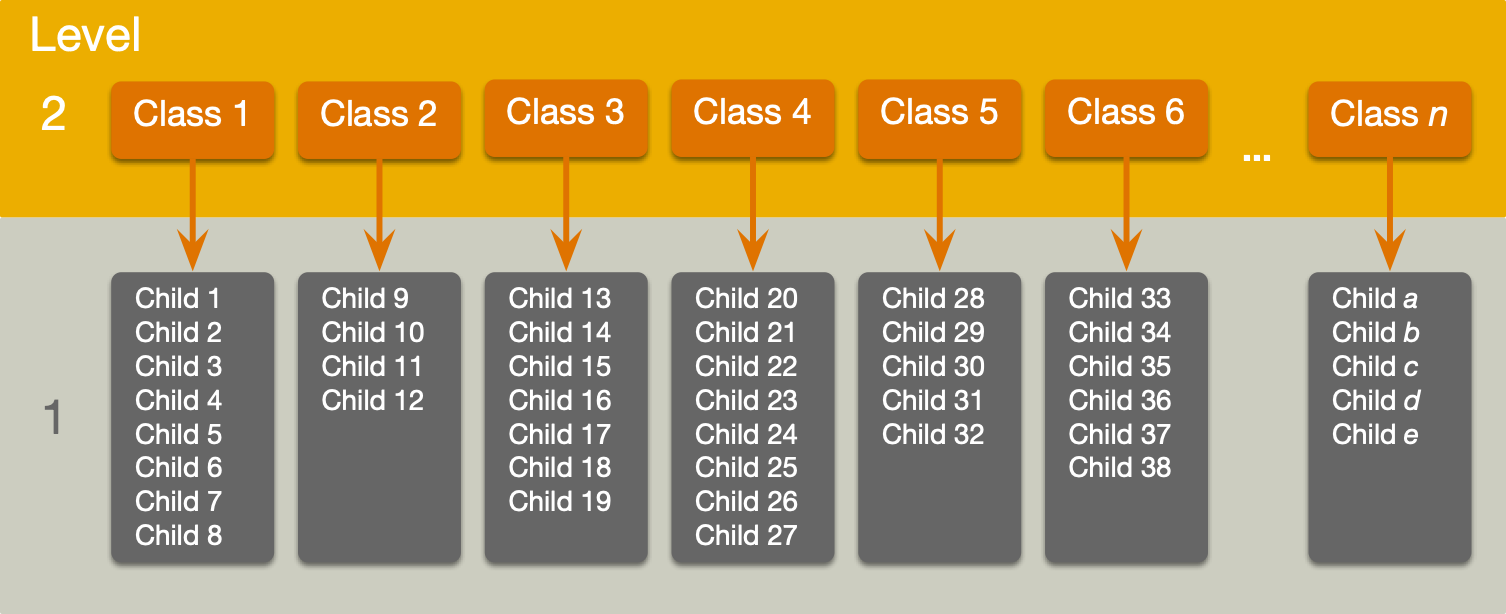
A three-level hierarchy
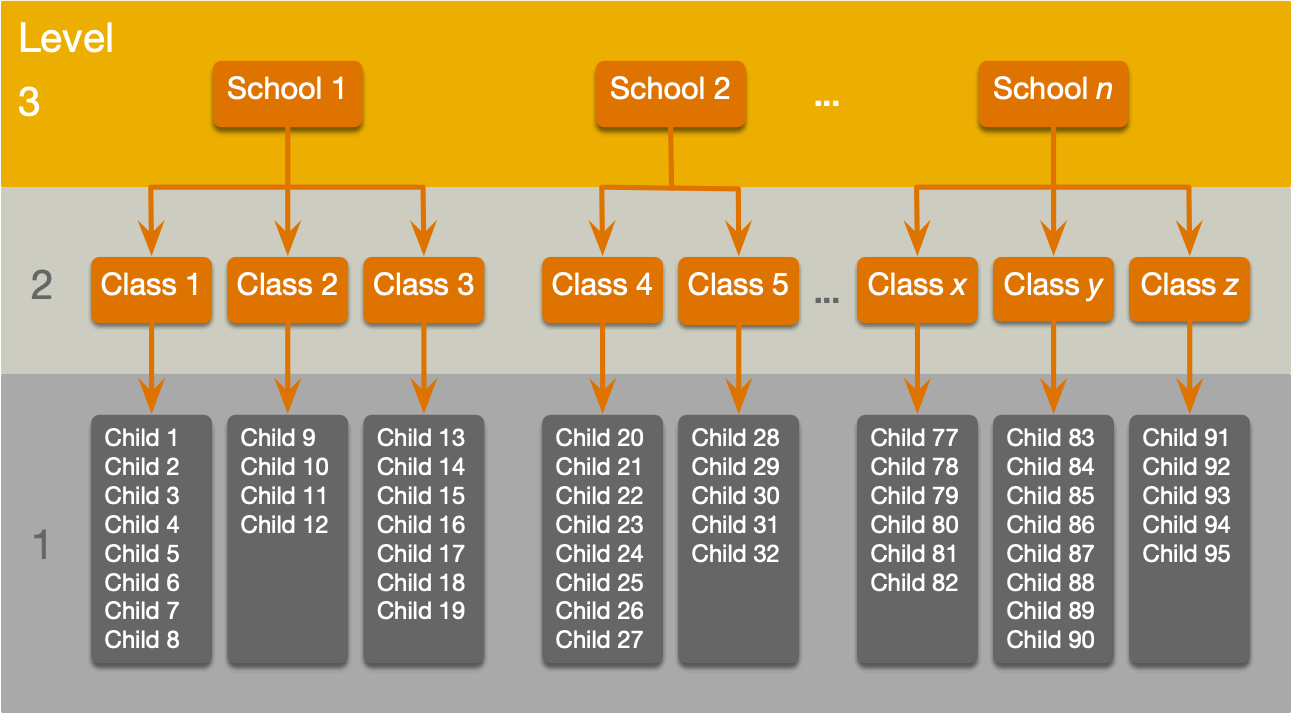
Why hierarchies matter
- Data from the same context will be more similar than data from different contexts
- Children in the same class will perform more similarly than children from different classes
- People treated in the same clinics should be more similar in response than those treated at different clinics
- Children in the same class will perform more similarly than children from different classes
- Lack of independence
- Violates the assumption of spherical errors (specifically, independence)
- Biases SEs, CIs and p-values


A surgical example
Research Questions
- Is quality of life after cosmetic surgery predicted by the length of time since surgery?
- Does this relationship depend on the reason for the surgery?
id: the participant’s participant codepost_qol: This is the outcome variable and it measures quality of life after cosmetic surgery.base_qol: We need to adjust our outcome for quality of life before the surgery.days: The number of days after surgery that post-surgery quality of life was measured.clinic: This variable specifies which of 21 clinics the person attended to have their surgery.reason: This variable specifies whether the person had surgery purely to change their appearance or because of a physical reason.
Let’s start with ….
\[ \begin{aligned} \text{QoL}_i &= \beta_0 + \beta_1\text{Days}_i + \varepsilon_i \\ \varepsilon_i &\sim N(0,\sigma^2) \end{aligned} \]
The surgery data hierarchy
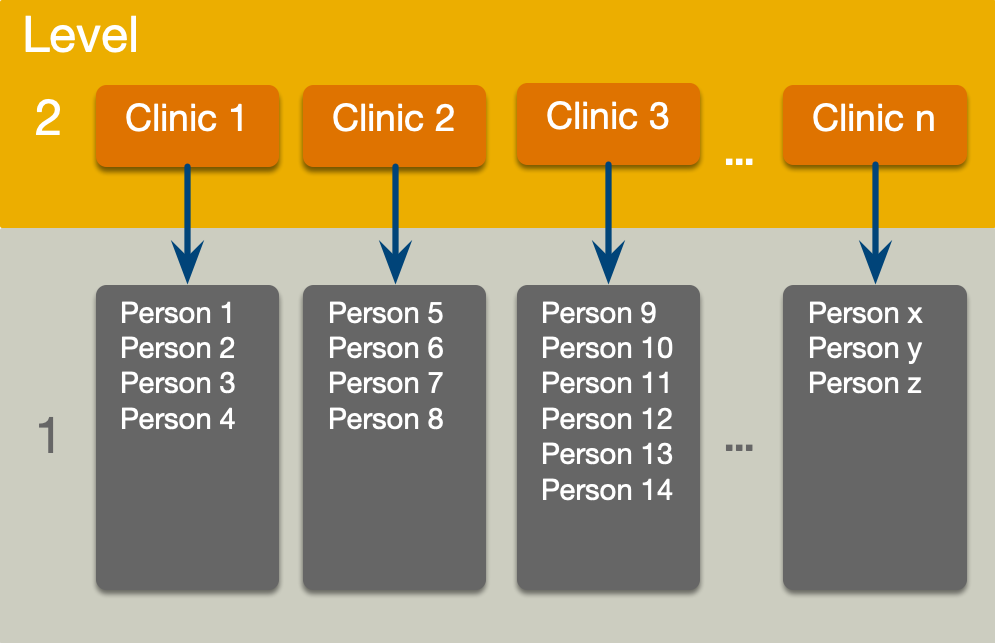
Fixed and random coefficients
- Intercepts and slopes can be fixed or random
- In OLS regression they are fixed
- Fixed coefficients
- Intercepts/slopes are assumed to be the same across different contexts (in this case clinics)
- Random coefficients
- Intercepts/slopes are allowed to vary across different contexts (in this case clinics)
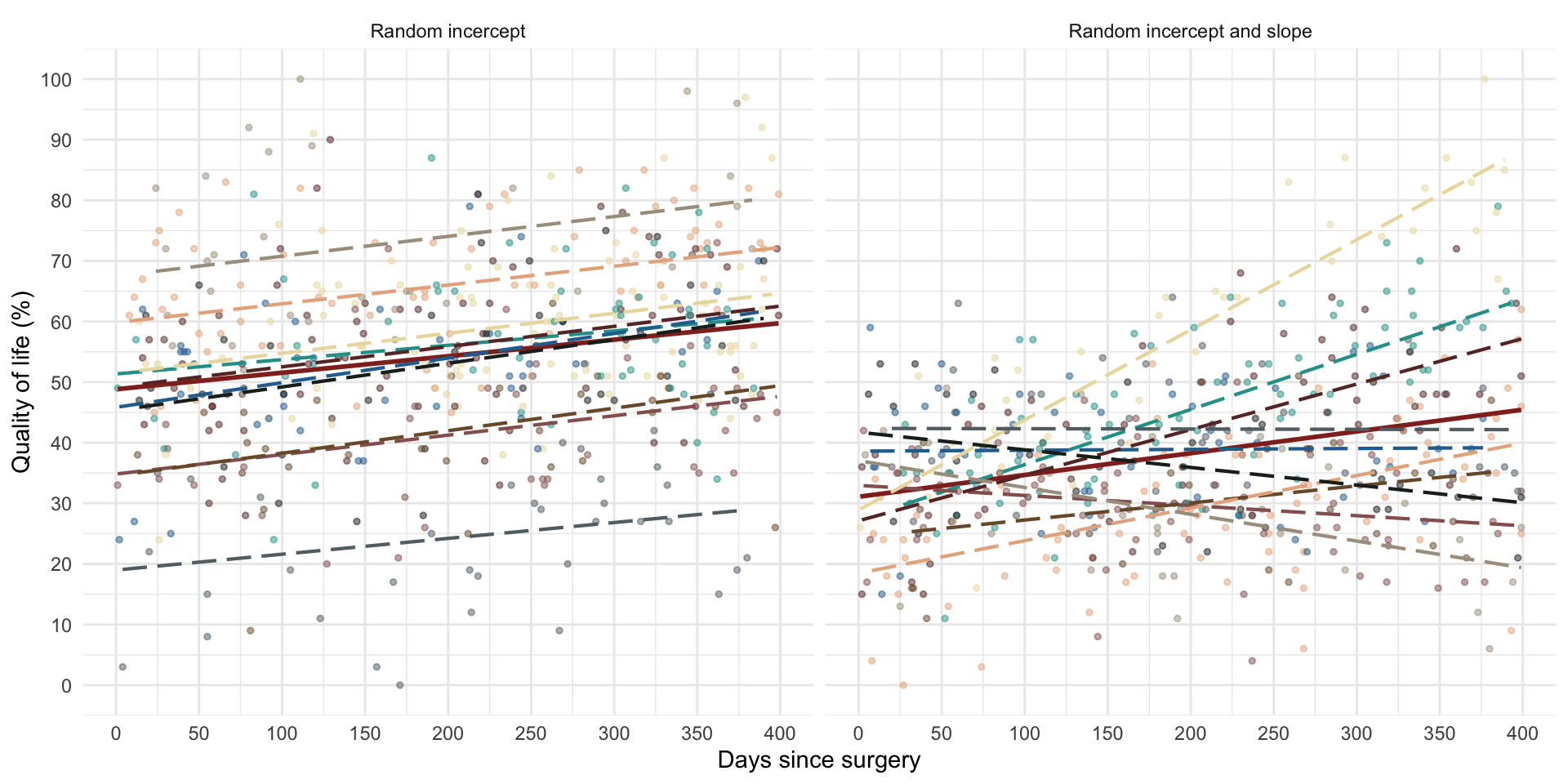
Random intercept
OLS model
\[ \begin{aligned} \text{QoL}_i &= \beta_0 + \beta_1\text{Days}_i + \varepsilon_i \\ \end{aligned} \]
Random intercept model (composite)
\[ \begin{aligned} \text{QoL}_{ij} &= (\beta_0 + u_{0j}) + \beta_1\text{Days}_{ij} + \varepsilon_{ij} \\ \text{QoL}_{ij} &= [\beta_0 + \beta_1\text{Days}_{ij}] + [u_{0j} + \varepsilon_{ij}] \\ \end{aligned} \]
Random intercept model (alternative)
\[ \begin{aligned} \text{QoL}_{ij} &= \beta_{0j} + \beta_1\text{Days}_{ij} + \varepsilon_{ij} \\ \beta_{0j} &= \beta_0 + u_{0j} \\ \end{aligned} \]
Random intercept
\[ \begin{aligned} \text{QoL}_{ij} &= [\beta_0 + \beta_1\text{Days}_{ij}] + [u_{0j} + \varepsilon_{ij}] \\ \end{aligned} \]
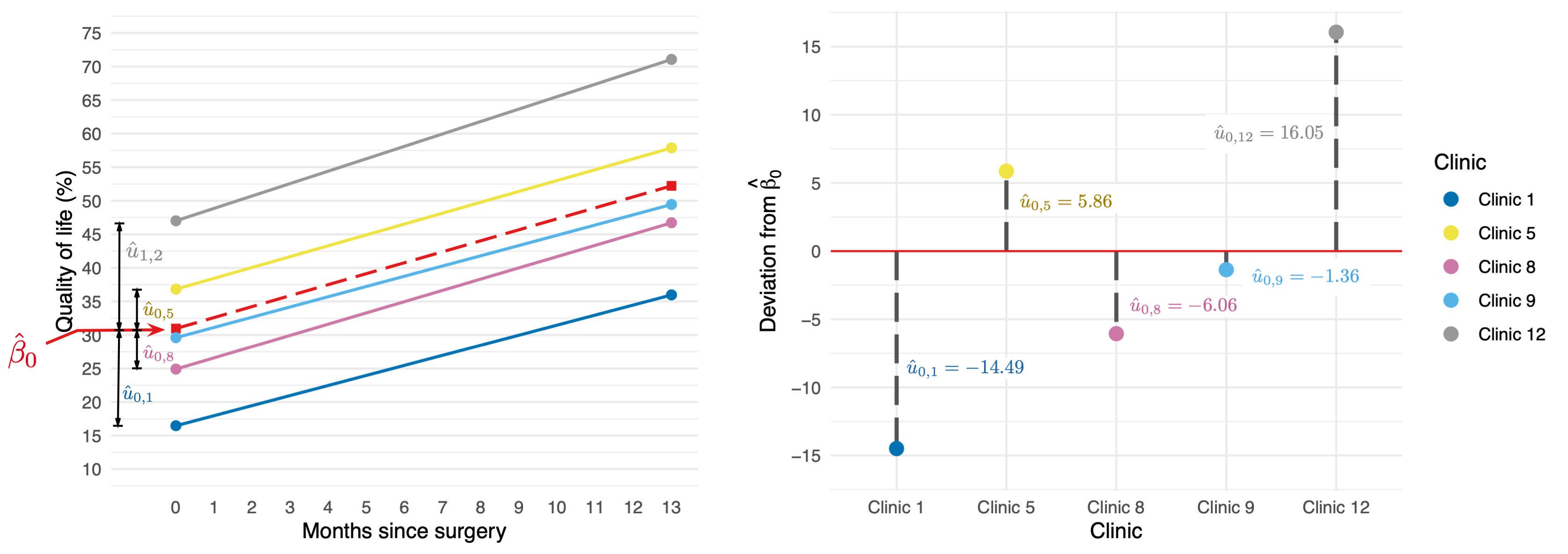
\[ \begin{aligned} u_0 \sim N(0, \sigma^2_{u_0}) \end{aligned} \]
Random slope
Random intercept model (composite)
\[ \begin{aligned} \text{QoL}_{ij} &= (\beta_0 + u_{0j}) + \beta_1\text{Days}_{ij} + \varepsilon_{ij} \\ \text{QoL}_{ij} &= [\beta_0 + \beta_1\text{Days}_{ij}] + [u_{0j} + \varepsilon_{ij}] \\ \end{aligned} \]
Random slope model (composite)
\[ \begin{aligned} \text{QoL}_{ij} &= (\beta_0 + u_{0j}) + (\beta_1 + u_{1j})\text{Days}_{ij} + \varepsilon_{ij} \\ \text{QoL}_{ij} &= [\beta_0 + \beta_1\text{Days}_{ij}] + [u_{0j} + u_{1j}\text{Days}_{ij} + \varepsilon_{ij}] \\ \end{aligned} \]
Random slope model (alternative)
\[ \begin{aligned} \text{QoL}_{ij} &= \beta_{0j} + u_{0j} + \beta_1\text{Days}_{ij} + \varepsilon_{ij} \\ \beta_{0j} &= \beta_0 + u_{0j} \\ \beta_{1j} &= \beta_1 + u_{1j} \\ \end{aligned} \]
Random slope
\[ \begin{aligned} \text{QoL}_{ij} &= [\beta_0 + \beta_1\text{Days}_{ij}] + [u_{0j} + u_{1j}\text{Days}_{ij} + \varepsilon_{ij}] \\ \end{aligned} \]
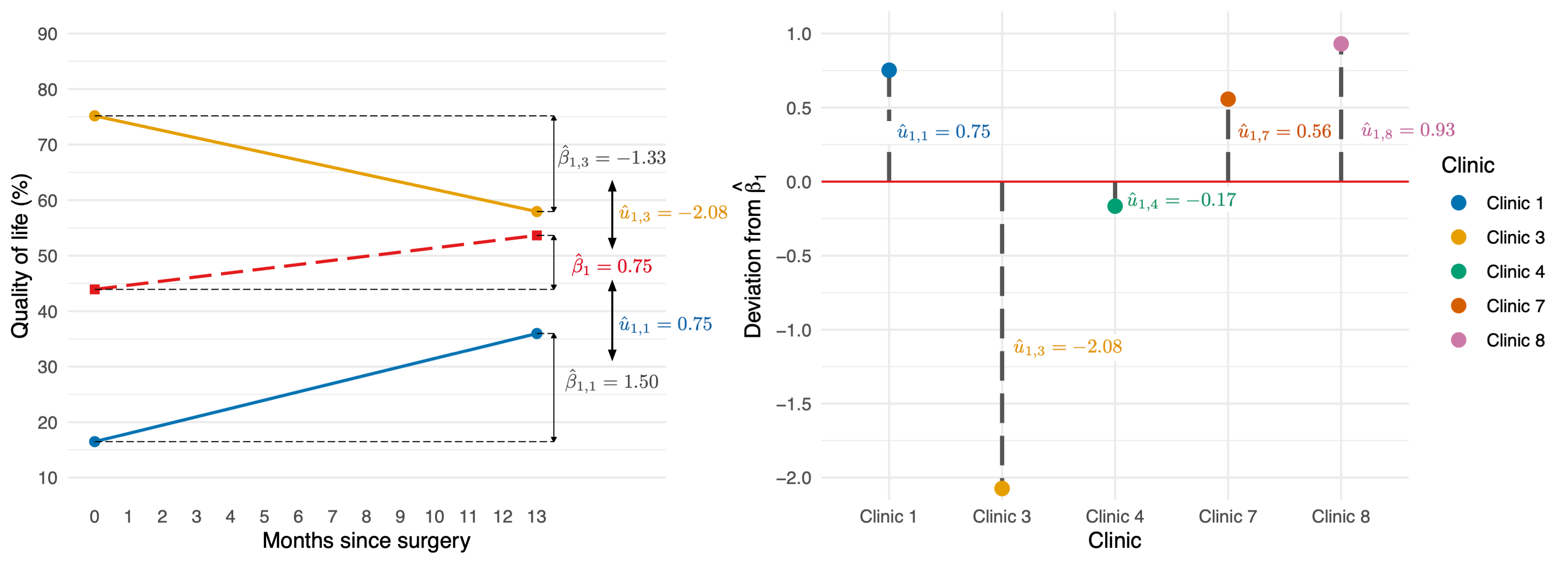
\[ \begin{aligned} u_1 \sim N(0, \sigma^2_{u_1}) \end{aligned} \]
The covariance structure of random effects
Random intercept and slope for 1 predictor
\[ \begin{aligned} \begin{bmatrix} u_0 \\ u_1 \end{bmatrix} \sim N\Bigg( \begin{bmatrix} 0 \\ 0 \end{bmatrix}, \begin{bmatrix} \sigma^2_{u_0} & \sigma_{u_0, u_1}\\ \sigma_{u_0, u_1} & \sigma^2_{u_1} \end{bmatrix} \Bigg) \end{aligned} \]
Random intercept and slope for several predictor
\[ \begin{aligned} \begin{bmatrix} u_0 \\ u_1 \\ \vdots \\ u_n \end{bmatrix} \sim N\begin{pmatrix}\begin{bmatrix} 0 \\ 0 \\ \vdots \\ 0 \end{bmatrix}, \begin{bmatrix} \sigma^2_{u_0} & \sigma_{u_0, u_1} &\dots & \sigma_{u_0, u_n}\\ \sigma_{u_0, u_1} & \sigma^2_{u_1} & \dots & \sigma_{u_1, u_n}\\ \vdots & \vdots & \ddots & \vdots\\ \sigma_{u_0, u_n} & \sigma^2_{u_1} & \dots & \sigma^2_{u_n}\\ \end{bmatrix} \end{pmatrix} \end{aligned} \]
Convergence
- As you include more random effects the number of parameters that need to be estimated from the data rapidly increases
- This increases the likelihood that the model won’t converge.
Level 1 errors

Normally distrubuted errors
\[ \begin{aligned} \varepsilon_{ij} &\sim N(0,\sigma^2) \end{aligned} \]
The covariance structure of level 1 errors
Spherical errors
\[ \begin{aligned} \Phi = \begin{bmatrix} \sigma^2_1 & 0 & 0 &\dots & 0\\ 0 & \sigma^2_2 & 0 & \dots & 0\\ 0 & 0 & \sigma^2_3 & \dots & 0\\ \vdots & \vdots & \ddots & \vdots\\ 0 & 0 & 0 & \dots & \sigma^2_n\\ \end{bmatrix} \end{aligned} \]
(Potential) Benefits of MLMs
- Modelling variability in effects across contexts
- Model the variability in intercepts
- Model the variability in slopes
- Model violations of the assumption of spherical errors
- Model differences in the variability of errors
- Model relationships between errors
- (Linear model for repeated observations – next two weeks!)
- Missing data
- MLMs (in general) cope with missing data
Model assumptions
- MLMs use maximum likelihood estimation not OLS
- Familiar assumptions
- Linearity and additivity
- Level 1 errors are normally distributed with mean of zero and constant variance (i.e. homoscedasticity)
- Independent errors (but we can model dependency)
- New assumptions
- Random effects (slopes and intercepts) are assumed to be normally distributed with mean of zero and constant variance (i.e. homoscedasticity)
Practical issues
Computing p-values
- There is no unifying method to compute p-values in multilevel models because the degrees of freedom of the test statistic are rarely known.
- df can be approximated (e.g., Satterthwaite and Kenward-Roger methods) but it’s unclear how good these approximations are for complex models/complex covariance structures.
Should effects be fixed or random?
- Three approaches
- Theory-driven
- Maximal model (Barr et al., 2013)
- Data-driven (include random effects that improve fit)
- Treat a predictor as a random effect if … (Bolker, 2015)
- You’re not interested in differences between the levels.
- You’re interested in quantifying the variability across levels of the variable.
- You’re interested in generalizing beyond the observed levels of the contextual variable.
- You have an unbalanced design.
- You have a categorical predictor that is not direct relevant to the hypothesis but for which you need to adjust (a nuisance variable).
The model we will fit
Composite form
\[ \begin{aligned} \text{QoL}_{ij} &= [\beta_0 + \beta_1\text{Days}_{ij} + \beta_2\text{Pre QoL}_{ij} + \beta_3\text{Reason}_{ij} + \beta_4\text{Days} \times \text{Reason}_{ij}] \\ &\quad + [u_{0j} + u_{1j}\text{Days}_{ij}+ \varepsilon_{ij}] \end{aligned} \]
Separate equations
\[ \begin{aligned} \text{QoL}_{ij} &= \beta_{0j} + \beta_{1j}\text{Days}_{ij} + \beta_2\text{Pre QoL}_{ij} + \beta_3\text{Reason}_{ij} + \beta_4\text{Days} \times \text{Reason}_{ij} + \varepsilon_{ij}\\ \beta_{0j} &= \beta_{0} + u_{0j} \\ \beta_{1j} &= \beta_{1} + u_{1j} \end{aligned} \]
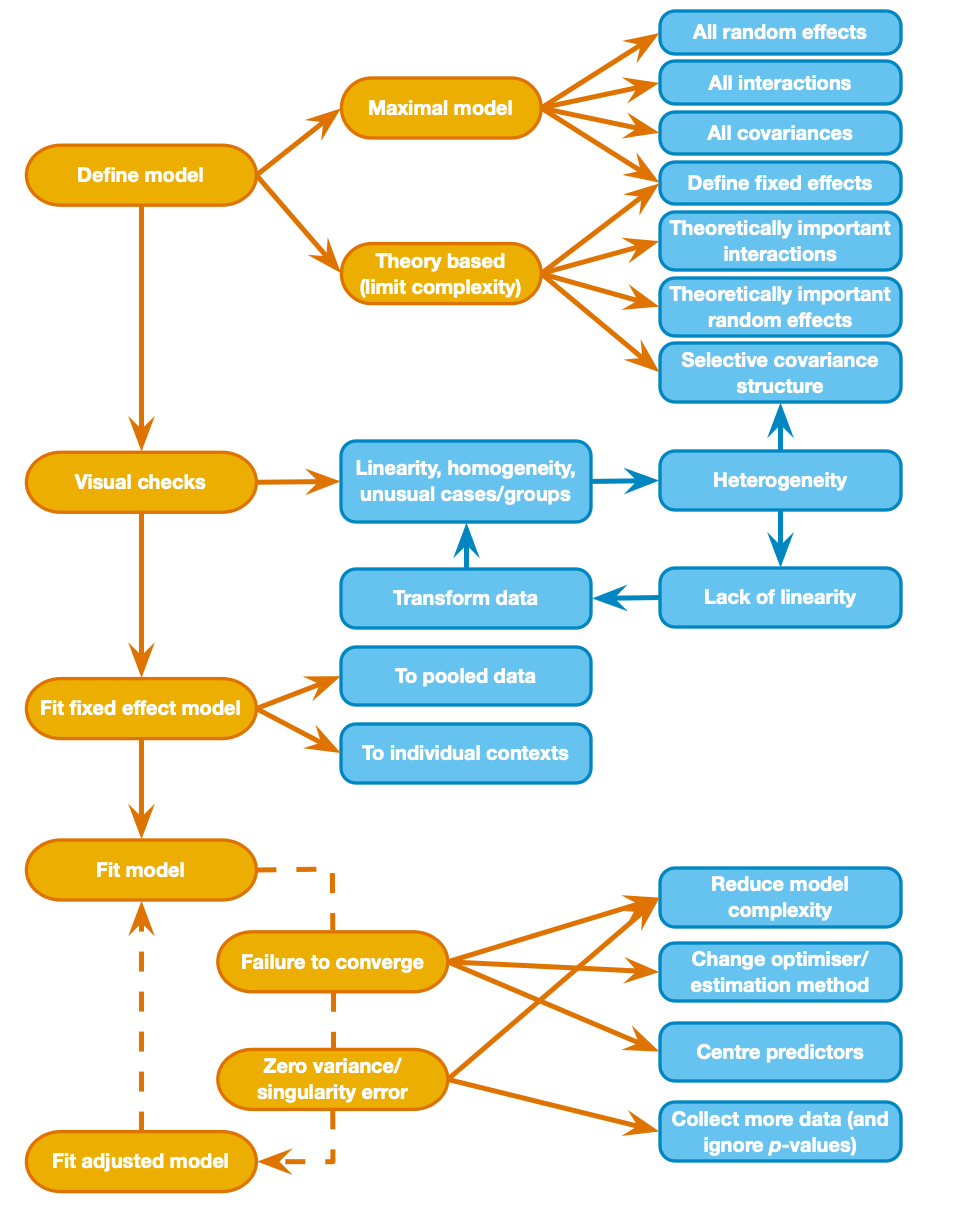
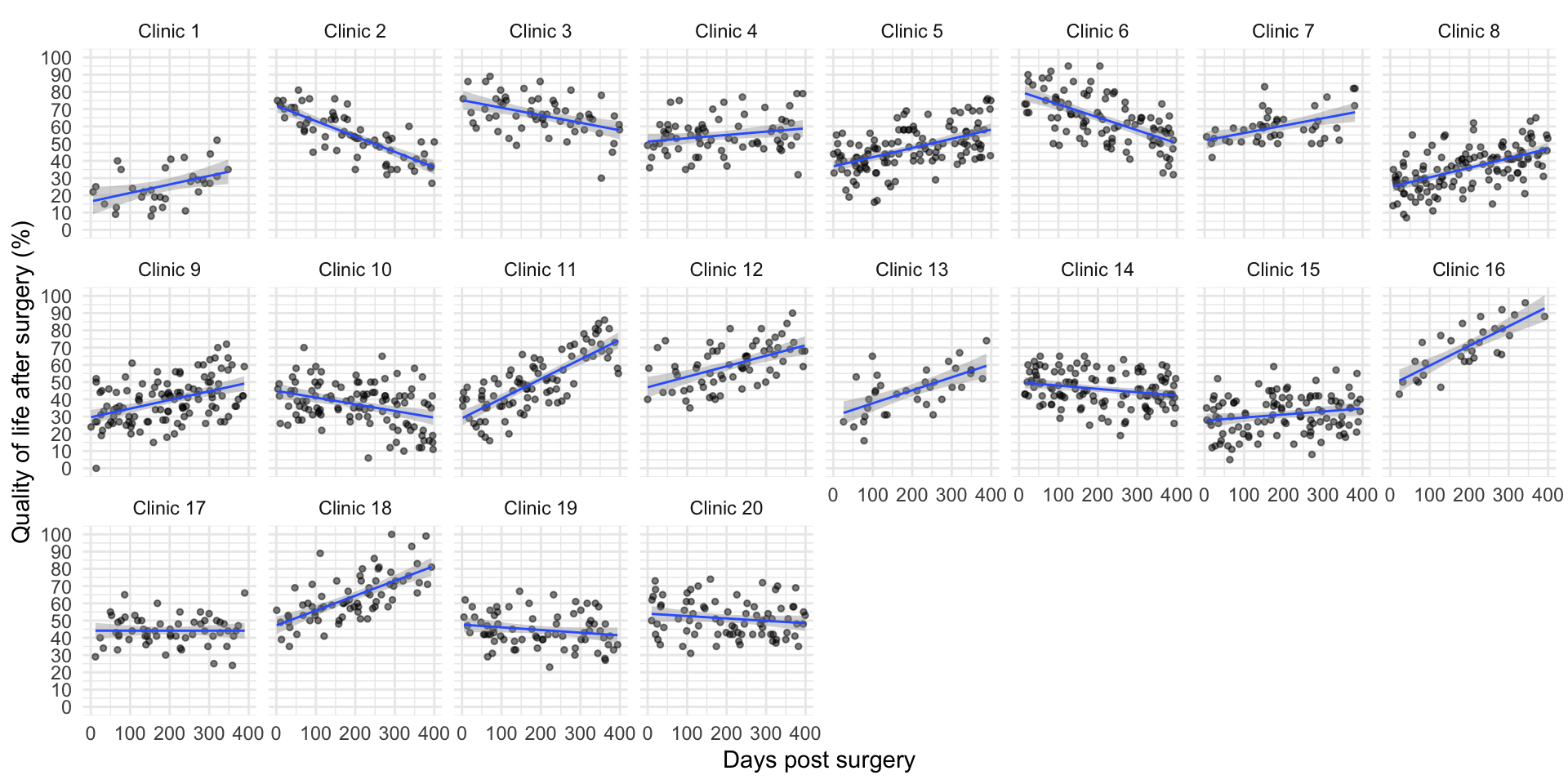
Visualise the data
Fitting a fixed-effect model on the pooled data
| term | estimate | std.error | statistic | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|
| (Intercept) | 24.75 | 2.06 | 12.03 | 0.00 | 20.72 | 28.79 |
| days | 0.01 | 0.00 | 2.07 | 0.04 | 0.00 | 0.02 |
| reasonPhysical reason | -2.48 | 1.62 | -1.53 | 0.13 | -5.65 | 0.69 |
| base_qol | 0.46 | 0.04 | 11.52 | 0.00 | 0.38 | 0.54 |
| days:reasonPhysical reason | 0.02 | 0.01 | 3.20 | 0.00 | 0.01 | 0.04 |
Fitting fixed-effect models within clinics
(Attempt to) fit the model
Convergence failure
- Warning: Model failed to converge with max|grad| = 6.14912 (tol = 0.002, component 1)
- Warning: Model is nearly unidentifiable: very large eigenvalue - Rescale variables?
cosmetic_mod |>
broom.mixed::tidy(conf.int = T) |>
dplyr::select(-c(effect, group)) |>
knitr::kable(digits = 3) |>
kableExtra::column_spec(4, background = "yellow")| term | estimate | std.error | statistic | df | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|---|
| (Intercept) | 25.081 | 3.599 | 6.969 | 33.008 | 0.000 | 17.759 | 32.404 |
| days | 0.016 | 0.012 | 1.283 | 23.452 | 0.212 | -0.010 | 0.041 |
| reasonPhysical reason | -1.799 | 0.979 | -1.838 | 1532.821 | 0.066 | -3.719 | 0.121 |
| base_qol | 0.470 | 0.024 | 19.523 | 1532.085 | 0.000 | 0.423 | 0.517 |
| days:reasonPhysical reason | 0.015 | 0.004 | 3.441 | 1532.380 | 0.001 | 0.006 | 0.023 |
| sd__(Intercept) | 15.069 | NA | NA | NA | NA | NA | NA |
| cor__(Intercept).days | -0.661 | NA | NA | NA | NA | NA | NA |
| sd__days | 0.054 | NA | NA | NA | NA | NA | NA |
| sd__Observation | 9.272 | NA | NA | NA | NA | NA | NA |
Fit the model again
| Sum Sq | Mean Sq | NumDF | DenDF | F value | Pr(>F) | |
|---|---|---|---|---|---|---|
| months | 275.41 | 275.41 | 1 | 19.03 | 3.21 | 0.09 |
| reason | 288.06 | 288.06 | 1 | 1535.47 | 3.36 | 0.07 |
| base_qol | 32785.53 | 32785.53 | 1 | 1534.92 | 382.66 | 0.00 |
| months:reason | 1014.54 | 1014.54 | 1 | 1535.12 | 11.84 | 0.00 |
| term | estimate | std.error | statistic | df | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|---|
| (Intercept) | 25.069 | 4.015 | 6.243 | 22.484 | 0.000 | 16.752 | 33.386 |
| months | 0.482 | 0.397 | 1.215 | 19.617 | 0.239 | -0.346 | 1.311 |
| reasonPhysical reason | -1.792 | 0.977 | -1.834 | 1535.469 | 0.067 | -3.709 | 0.125 |
| base_qol | 0.470 | 0.024 | 19.562 | 1534.925 | 0.000 | 0.423 | 0.517 |
| months:reasonPhysical reason | 0.447 | 0.130 | 3.441 | 1535.124 | 0.001 | 0.192 | 0.703 |
Interpretation
| term | estimate | std.error | statistic | df | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|---|
| (Intercept) | 25.069 | 4.015 | 6.243 | 22.484 | 0.000 | 16.752 | 33.386 |
| months | 0.482 | 0.397 | 1.215 | 19.617 | 0.239 | -0.346 | 1.311 |
| reasonPhysical reason | -1.792 | 0.977 | -1.834 | 1535.469 | 0.067 | -3.709 | 0.125 |
| base_qol | 0.470 | 0.024 | 19.562 | 1534.925 | 0.000 | 0.423 | 0.517 |
| months:reasonPhysical reason | 0.447 | 0.130 | 3.441 | 1535.124 | 0.001 | 0.192 | 0.703 |
Write-up
- The overall effect of time on quality of life was small and non-significant, \(\beta\) = 0.48 [−0.35, 1.31], t(19.62) = 1.225, p = 0.239.
| term | estimate | std.error | statistic | df | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|---|
| (Intercept) | 25.069 | 4.015 | 6.243 | 22.484 | 0.000 | 16.752 | 33.386 |
| months | 0.482 | 0.397 | 1.215 | 19.617 | 0.239 | -0.346 | 1.311 |
| reasonPhysical reason | -1.792 | 0.977 | -1.834 | 1535.469 | 0.067 | -3.709 | 0.125 |
| base_qol | 0.470 | 0.024 | 19.562 | 1534.925 | 0.000 | 0.423 | 0.517 |
| months:reasonPhysical reason | 0.447 | 0.130 | 3.441 | 1535.124 | 0.001 | 0.192 | 0.703 |
Write-up
- The effect of the reason for surgery was also small and non-significant, \(\beta\) = −1.79 [−3.71, 0.12], t(1535.47) = −1.83, p = 0.067.
- With other variables held constant the difference in quality of life in those seeking surgery for physical reasons was 1.79 lower (on the 100-point scale) than those seeking it for cosmetic reasons.
| term | estimate | std.error | statistic | df | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|---|
| (Intercept) | 25.069 | 4.015 | 6.243 | 22.484 | 0.000 | 16.752 | 33.386 |
| months | 0.482 | 0.397 | 1.215 | 19.617 | 0.239 | -0.346 | 1.311 |
| reasonPhysical reason | -1.792 | 0.977 | -1.834 | 1535.469 | 0.067 | -3.709 | 0.125 |
| base_qol | 0.470 | 0.024 | 19.562 | 1534.925 | 0.000 | 0.423 | 0.517 |
| months:reasonPhysical reason | 0.447 | 0.130 | 3.441 | 1535.124 | 0.001 | 0.192 | 0.703 |
Write-up
- The effect of baseline quality of life on post-surgery quality of life was substantial and significant, \(\beta\) = 0.47 [0.42, 0.52], t(1534.92) = 19.56, p < 0.001.
- With other variables held constant, for every unit increase in baseline quality of life there is a half unit increase in post-surgery quality of life, which is a fairly strong relationship.
| term | estimate | std.error | statistic | df | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|---|
| (Intercept) | 25.069 | 4.015 | 6.243 | 22.484 | 0.000 | 16.752 | 33.386 |
| months | 0.482 | 0.397 | 1.215 | 19.617 | 0.239 | -0.346 | 1.311 |
| reasonPhysical reason | -1.792 | 0.977 | -1.834 | 1535.469 | 0.067 | -3.709 | 0.125 |
| base_qol | 0.470 | 0.024 | 19.562 | 1534.925 | 0.000 | 0.423 | 0.517 |
| months:reasonPhysical reason | 0.447 | 0.130 | 3.441 | 1535.124 | 0.001 | 0.192 | 0.703 |
Write-up
- The combined effect of months and reason on post-surgery quality of life was significant, \(\beta\) = 0.45 [0.19, 0.70], t(1535.12) = 3.44, p = 0.001.
- The change over time of quality of life is 0.45 bigger in those having surgery for physical reasons than in those having it for cosmetic reasons.
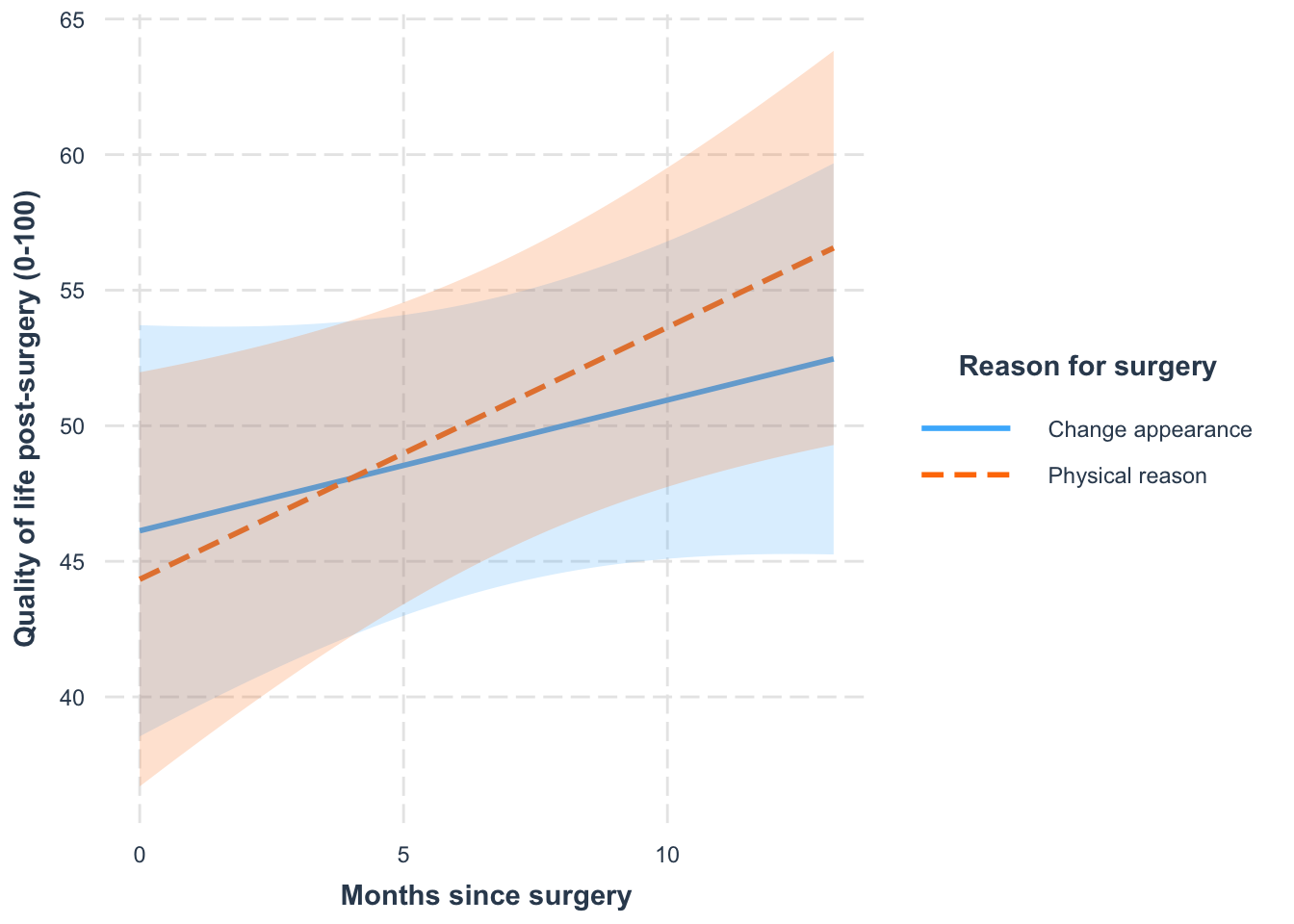
Simple slopes
| reason | Coefficient | SE | CI | CI_low | CI_high | t | df_error | p |
|---|---|---|---|---|---|---|---|---|
| Change appearance | 0.482 | 0.397 | 0.95 | -0.346 | 1.311 | 1.215 | 19.623 | 0.239 |
| Physical reason | 0.930 | 0.401 | 0.95 | 0.094 | 1.765 | 2.316 | 20.558 | 0.031 |
Write-up
- For those who had surgery to change their appearance, their quality of life increased over time but not significantly so, \(\beta\) = 0.48 [−0.35, 1.31], t(19.62) = 1.22, p = 0.239. Quality of life increased by 5.76 units (on the 100-point scale) per year.
- For those who had surgery to help with a physical problem, their quality of life significantly increased over time, \(\beta\) = 0.93 [0.09, 1.77], t(20.56) = 2.32, p = 0.031. Quality of life increased by 11.16 units (on the 100-point scale) per year.
Summary
- Data can be hierarchical and this hierarchical structure can be important.
- The OLS linear model simply ignores the hierarchy.
- Hierarchical models are just a fancy linear model in which you estimate the variability in the slopes and intercepts within contexts
- i.e. slopes and intercepts can be random variables (allowed to vary) rather than fixed (assumed to be equal in different situations).
- MLMs are a world of pain